Research Article - Imaging in Medicine (2026) Volume 17, Issue 2
Magnetic Resonance Imaging Image Analysis Using Diffusion Weighting
- Corresponding Author:
- Flaviana
Department of Physics, Parahyangan Catholic University, Bandung, Indonesia
E-mail: flaviana@unpar.ac.id
Received: 12-September-2024, Manuscript No. FMIM-24-147840; Editor assigned: 16-September-2024, PreQC No. FMIM-24-147840 (PQ); Reviewed: 30-September-2024, QC No. FMIM-24-147840; Revised: 15-Feb-2025, Manuscript No. FMIM-24-147840 (R); Published: 22- Feb-2025, DOI: 10.47532/1755-5191.2025.17(1).269-276
Abstract
The brain is one of the largest organs in the human body and plays a major role in visual processing, hearing, memory, body movement control and so on. Microscopically, the human brain can be described as the random movement of water molecules within the brain tissue. By making further observations of these molecular movements, tissue boundaries within the brain can be visualized. In the field of medicine, several visual diagnostic tools are known to detect and evaluate the brain's response and function. Magnetic Resonance Imaging (MRI) is a medical imaging technique that provides important information about the soft tissue anatomy of the human body. By scanning the brain, doctors can diagnose if there are abnormalities in the brain. MRI image processing and analysis is an important part of the MRI modality, which is to improve image quality and extract information from MRI image data in an efficient and accurate manner. Diffusion-Weighted Imaging (DWI) provides significant structural information of the brain at the cellular level, emphasizing aspects of brain pathophysiology. In theory, DWI is based on the motion of water molecules so that it can describe tissue microstructure, and the opportunity to perform tumor characterization that can provide a new perspective for clinical experts in developing brain tumor treatment. The development of image processing techniques in the medical field today allows for more in-depth analysis of MRI scanning images that have been used in hospitals. This study aims to classify area groups in Diffusion Weighting (DW) MRI phase images, make statistical observations of the area groups, and evaluate the extent to which the image classification has been effectively carried out.
Keywords
Diffusion weighting • Magnetic resonance imaging • Mixture model • Maximum likelihood • Expectation maximization
Introduction
The human brain is one of the most complex organs because it consists of more than 100 billion nerve cells that are connected to each other to transmit information through electrical or chemical signals [1]. Based on the parts of the lobes in the brain, the brain has a major role in visual, auditory, and tactile processing systems. In addition, the brain's functions are also related to memory, body movement control and so on [2].
In the field of medicine, several visual diagnostic tools are known to detect and evaluate the brain's responsiveness and functionality. Magnetic Resonance Imaging (MRI) is a medical imaging technique that provides important information about the anatomy of the body, especially the soft tissues of the human body [3]. By scanning the brain, doctors can diagnose if there is any abnormality in the brain.
Microscopically, the human brain can be described as the movement of water molecules in the brain tissue. The movement of water molecules influenced by thermal energy causes the molecules to diffuse in the body. The movement is restricted by other surrounding body tissues. By making further observations of this molecular movement, the boundaries of the tissues in the brain can be visualized.
The particles in a fluid are forced to move in a random motion that is often referred to as Brownian motion. The result of this random motion is diffusion. However, the diffusion motion of molecules in body tissues does not occur freely, but due to interactions with large molecules, membranes, fibers, and so on. Thus, the diffusion distance that occurs is relatively reduced when compared to the random motion of molecules in water freely. Statistical observation of the distribution of molecular motion provides unique information about the structural features and geometric arrangement of brain nerve tissue on a microscopic scale. MRI can thus be used to visually quantify the diffusion characteristics of the tissue [4].
MRI image processing and analysis is an important part of the MRI modality, which is to improve image quality and extract information from MRI image data in an efficient and accurate manner. However, conventional MRI examination has several limitations in detecting tumors, such as detection of tumor spread and tumor classification. The process of making a diagnosis becomes more accurate if the spread of tumor cells can be detected. Likewise, if precise tumor classification can be performed, the diagnosis process becomes more efficient [5].
When a radiofrequency pulse is applied during MRI scanning, the protons rotate at different rates depending on the strength, duration, and direction, which is called a gradient. By applying a gradient of equal to opposite magnitude, the protons are refocused. Thus, information on how many proton spins have moved or diffused in a given time can be obtained.
In Diffusion-Weighted (DW) MRI, what is measured is the relaxation (dephasing) of proton spins when given a certain magnetic field (gradient). When a patient is inserted in an MRI scanner that has a homogeneous magnetic field, the spin of the nucleus will move along the direction of the static magnetic field. The longer and stronger the gradient pulse, the greater the change in direction of the molecules. The greater the signal attenuation and loss of coherence due to macroscopic motion.
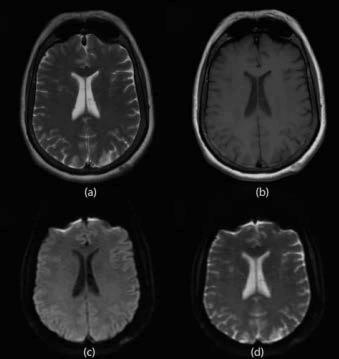
The brightness of each voxel in the image is related to the DW intensity, which is the sum of the diffusion weights or gradients. For example, water-like tissues (e.g. cerebrospinal fluid, CSF) that have high mobility protons will appear lower in intensity, while more static or dense tissues will give a stronger signal. Thus, DW contrast behaves like T1-weighting, or opposite to T2- weighting (Figure 1).
Figure 1. MRI images (A) T2, (B) T1, (C) DW, (D) ADC.
The development of image processing techniques in the medical field today allows for more in-depth analysis of MRI scanning images that have been used in hospitals. Analysis techniques that continue to develop are needed to support medical technology. This research aims to study the processing and analysis of brain MRI images through digital image samples to provide new information, ideas and perspectives related to the development of segmentation and image analysis methods. The purpose of image segmentation and analysis is to classify area groups in Diffusion Weighting (DW) MRI phase images, make statistical observations of the area groups, and evaluate the extent to which the image classification has been effectively carried out.
Materials and Methods
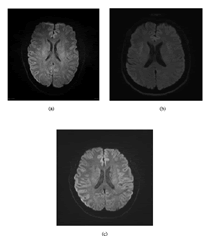
Diffusion-Weighted Imaging (DWI) provides significant structural information of the brain at the cellular level, emphasizing aspects of brain pathophysiology. In theory, DWI is based on the motion of water molecules, which can describe the microstructure of tissues, and the opportunity to perform tumor characterization, which can provide new perspectives for clinical experts in developing brain tumor treatments (Figure 2).
Figure 2. (A) DW MRI image of brain N1, (B) DW MRI image of brain N2, (C) DW MRI image of brain N3.
Previous studies have been conducted related to the processing of MRI images of tumors in the head and neck using MR DWI with a high b-value [6]. This technique is used in detecting tumors that reappear in the head and neck. Another research was developed by Villalon-Reina in 2016 which applied segmentation algorithms in the form of Bayes formula approach, Maximum A Posteriori, and expextation maximization on T1 phase MRI images [7].
The study began by compiling 3 samples of axial cross-sectional brain DW MRI images called N1, N2, N3, respectively. The data was taken from the open source www.radiopaedia.org. After the library materials (physical or digital) as well as image samples have been collected, the researchers discussed plans for image processing and analysis techniques that will be applied to the MRI images.
As explained in the previous section, the brightness of each voxel in a DW MRI image is its intensity, which is the sum of the diffusion weights or gradients. Water-like tissues that have protons with high mobility will appear lower in intensity, while tissues that are more static or dense will give a stronger signal or intensity.
In this study, the segmentation method applied to DW MRI images of the brain uses mixture modeling. With this approach, it is assumed that the pixel values of the MRI image come from varying densities (multimodal). The mode value contained in the image histogram indicates that there are several pixel intensity values, where the number of pixels containing these intensities is relatively large. The mode values that appear on the histogram can be interpreted as belonging to certain areas or types of tissue in the body. By applying the mixture model, we want to observe its effect on image contrast, as well as the extent to which the results are effective in classifying certain areas in DW MRI images of the brain.
Next, the probability density function (pdf ) of the pixel values is modeled. The model is then used to find the probability of pixel values being in different classes or areas [8].
Firstly, the image histogram information can provide a great overview of the probability density of pixel values. The image is processed in grayscale data type with a value range of 0 to 255. Furthermore, the pixel value x and f(x) is the probability of getting that pixel value. If there are k modes or classes, then the value of x can come from the k density with different probability values and is shown in Equation (1):

The equation can also be written in another form as shown in Equation (2):

Probability Pr (kelas i) rewritten into weights wi.
Furthermore, one of the practical ways used in estimating the unknown parameters of the mixture model can be chosen by increasing the maximum likelihood function through Equation (3), θ unknown parameters of the mixture model, can be chosen by enlarging the maximum likelihood function through Equation (3):
θML=argθ maxL(θ) (3)
Where L(θ) is the maximum logarithm of the likelihood function (Equation (4)) [9].

One method that is often used to optimize this mixed model is by using Expectation Maximization (EM). EM is an iterative method to optimize a likelihood function when there is some missing information. In this case, the missing information is the pixel values. The expectation step (Equation (5)) is performed by calculating the conditional pdf class using the estimated value of the pixel. θ and determining the posterior probability, xj:

Then calculate the new weight value, wi or prior probability by averaging the posterior probabilities for each class using the Bayes formula in Equation (6):

Furthermore, the Expectation Maximization (EM) step is carried out by updating the parameters θ by enlarging the likelihood function (Equation (7)), where μ Ìi is the mean vector and Σ Ìi is the covariance matrix.

Then repeating the expectation and maximization steps (iteration) until the relative change in the mixture parameter estimates. Up to this point, the process is often sufficient to determine the number of classes based on the ML-EM step pdf. But in some other cases, sometimes the determination of the number of classes in image classification cannot be confirmed. Therefore, in the final stage, a Bayesian information creation, BIC (equation (8)) quality fit test is performed to determine the optimal number of mixture component models.
BIC =-2L(θ)+Mlog(n) (8)
Where L(θ) is the maximum log-likelihood function, M is the number of model parameters, and n represents the number of pixels. From the equation, it can be seen that the value of -2L(θ) decreases when M increases [4]. This computational process was carried out using Matlab 2016a software.
Results and Discussion
From the process of identifying pixel values pdf, it can be used to find the probability that certain pixel values can be classified into a certain number of classes or components. From the three sample brain DW MRI images used, N1, N2, and N3, respectively, the optimal number of classes in the image can be seen using Equation (2) through Equation (7). In addition, several statistical values such as weight, mean, and standard deviation of the image classification process were calculated after iterating (Table 1).
| Sample | Component (class) | |||
| 1 | 2 | 3 | 4 | |
| N1 | ||||
| Weight | 0,49 | 0,18 | 0,32 | 0,01 |
| Mean | 1,84 | 9,51 | 83,49 | 236 |
| Standard deviation | 2,17 | 5,03 | 25,98 | 0 |
| Number of iterations=39 | ||||
| N2 | ||||
| Weight | 0,42 | 0,25 | 0,34 | |
| Mean | 8 | 14,11 | 49,28 | |
| Standard deviation | 0 | 5,55 | 6,46 | |
| Number of iterations=12 | ||||
| N3 | ||||
| Weight | 0,44 | 0,22 | 0,12 | 0,22 |
| Mean | 28,89 | 33,61 | 98,7 | 114,19 |
| Standard deviation | 1,81 | 3,71 | 33,27 | 15,15 |
| Number of iterations=200 | ||||
TABLE 1.The statistical results of proposed approach.
In the process of classifying the N1 image, it is shown that the optimal number of classes obtained is 4 classes with a mean ± SD value of pixel intensity of classes 1 to 4 respectively of 1.84 ± 2.17; 9.51 ± 5.03; 83.49 ± 25.98; 236 ± 0 (gray scale value range 0-255). These results are after 39 iterations of the ML-EM process. It is indeed quite obvious visually the large variation in pixel intensity from dark to light. In addition, it can also be observed that from the classification of the 4 classes in the N1 image, respectively, the largest to smallest weights are the 1st; 3rd; 2nd; 4th classes.
Likewise, the number of classes of image N3 is obtained as 4 classes, after going through the iteration process 200 times. Visually, it can also be seen that the N3 image has a fairly large variation in color brightness, but the mean pixels of the 1st and 2nd classes are larger when compared to the N1 image (the background of the N3 image does not look so dark). Classifying the 4 classes in the N3 image, respectively, which have the largest weight to the smallest is the 1st class; the 2nd and 3rd classes have the same weight; the 4th class.
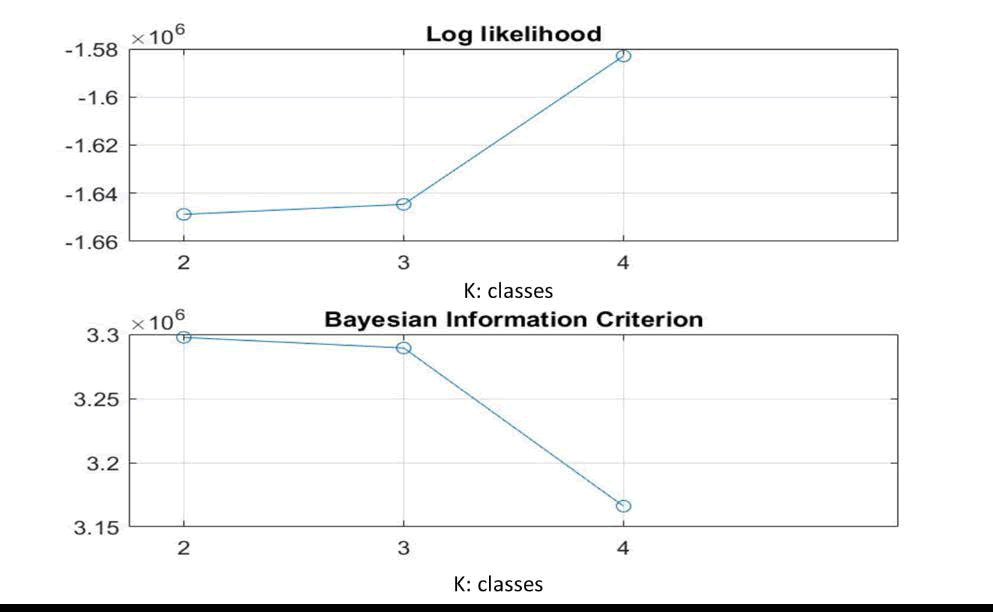
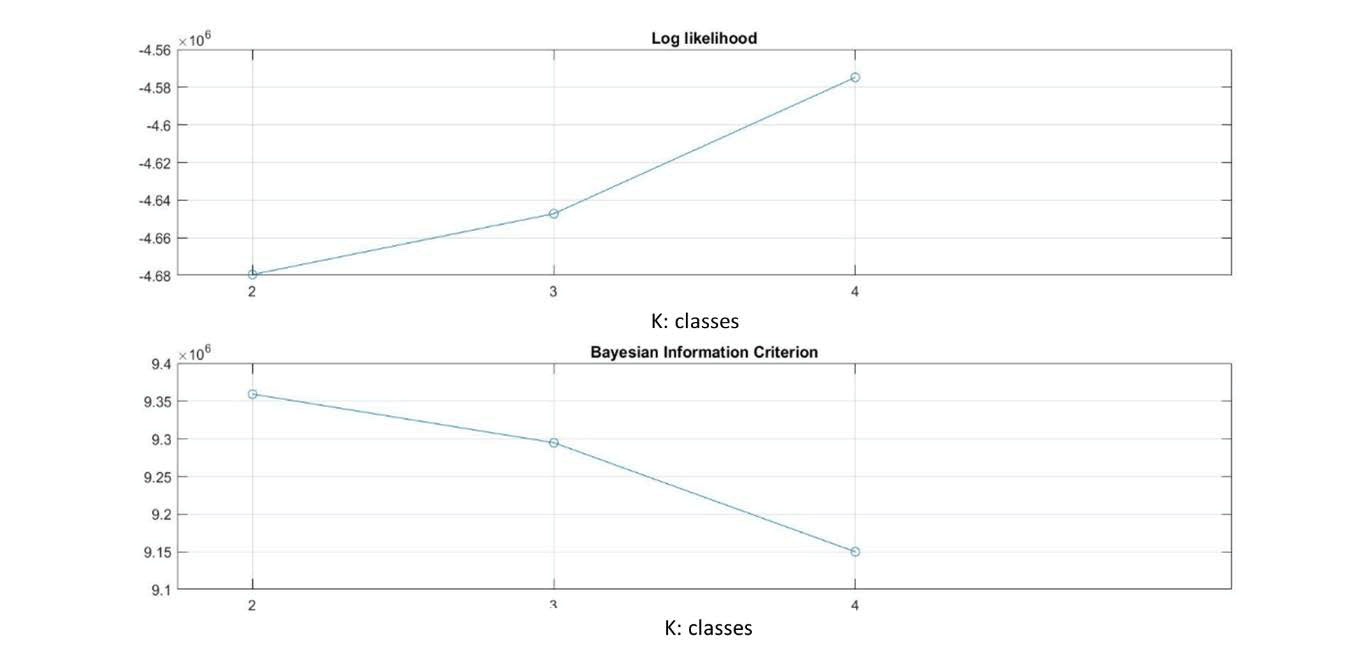
In the N2 image, the optimal number of classes obtained after 12 iterations is only 3 classes with mean ± SD values of pixel intensity of classes 1 to 3 respectively of 8 ± 0; 14.11 ± 5.55; 49.28 ± 6.46. The N2 image also visually does not appear to have a large variation in color brightness. In addition, the N1 image that has the largest to smallest weight is the 1st; 3rd; 2nd class, respectively. Furthermore, the results of classifying MRI images using the ML-EM mixture model are also shown through the maximum log-likelihood function graph (upper part of Figures 3-5). In addition, the results of the quality fit test using BIC are shown in Figures 3-5 at the bottom. The graph in Figure 3 below shows that the number of classes in the N1 image using the ML parameter model is 4 classes. While the BIC test results show the minimum in the 4th class. This is in accordance with Equation (8), where the minimum estimated missing pixel information indicates the most optimal ML model.
Figure 3. (top) Maximum likelihood (ML) graph, (b) BIC quality fit test of N1 image.
The result of the number of similar classes in Table 1 is shown in Figure 4. Image N2 has an optimal value of 3 classes, as well as the minimum BIC value.
Figure 4. (top) Maximum likelihood (ML) graph, (b) BIC quality fit test of N2 image.
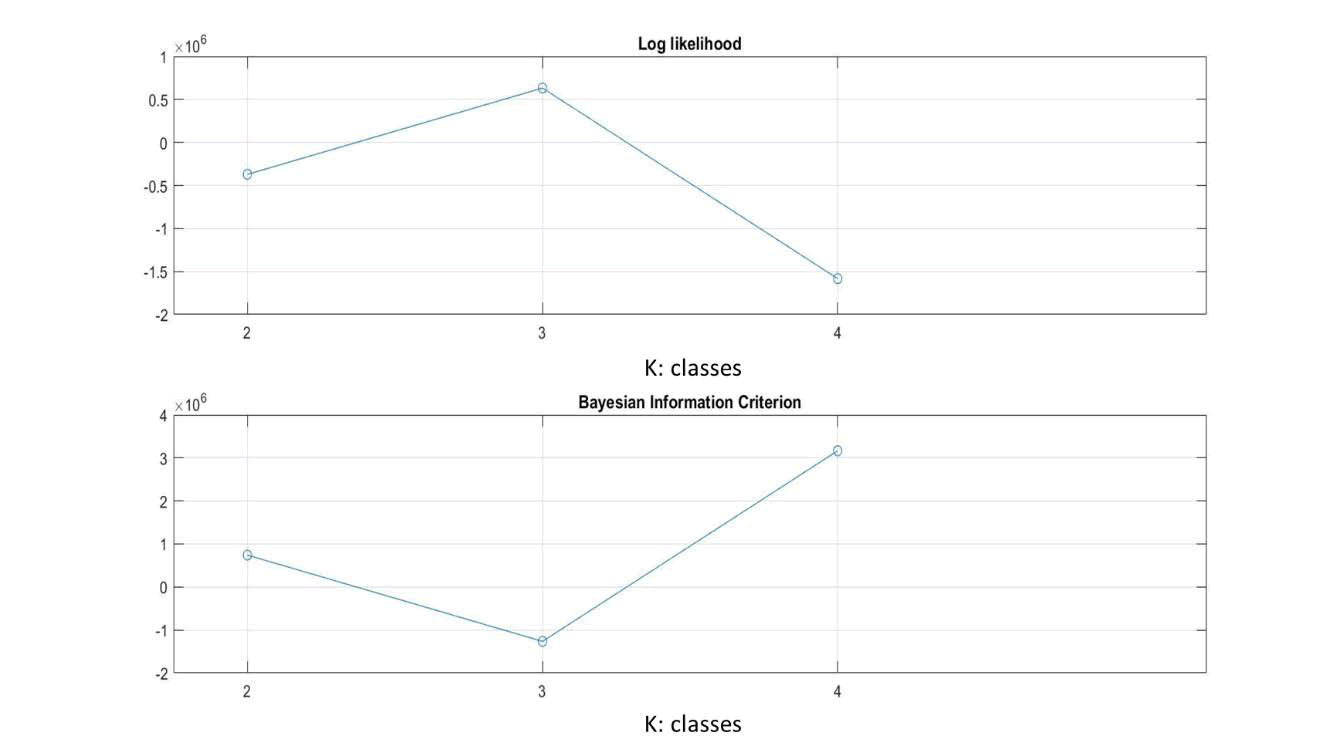
Figure 5. (top) Maximum likelihood (ML) graph, (bottom) BIC quality fit test of N3 image.
Figure 5 (top) shows that the optimal number of classes in the N3 image using the ML parameter model is 4 classes. While using the BIC test (Figure 5 below), it can be seen that the minimum value is at K=4.
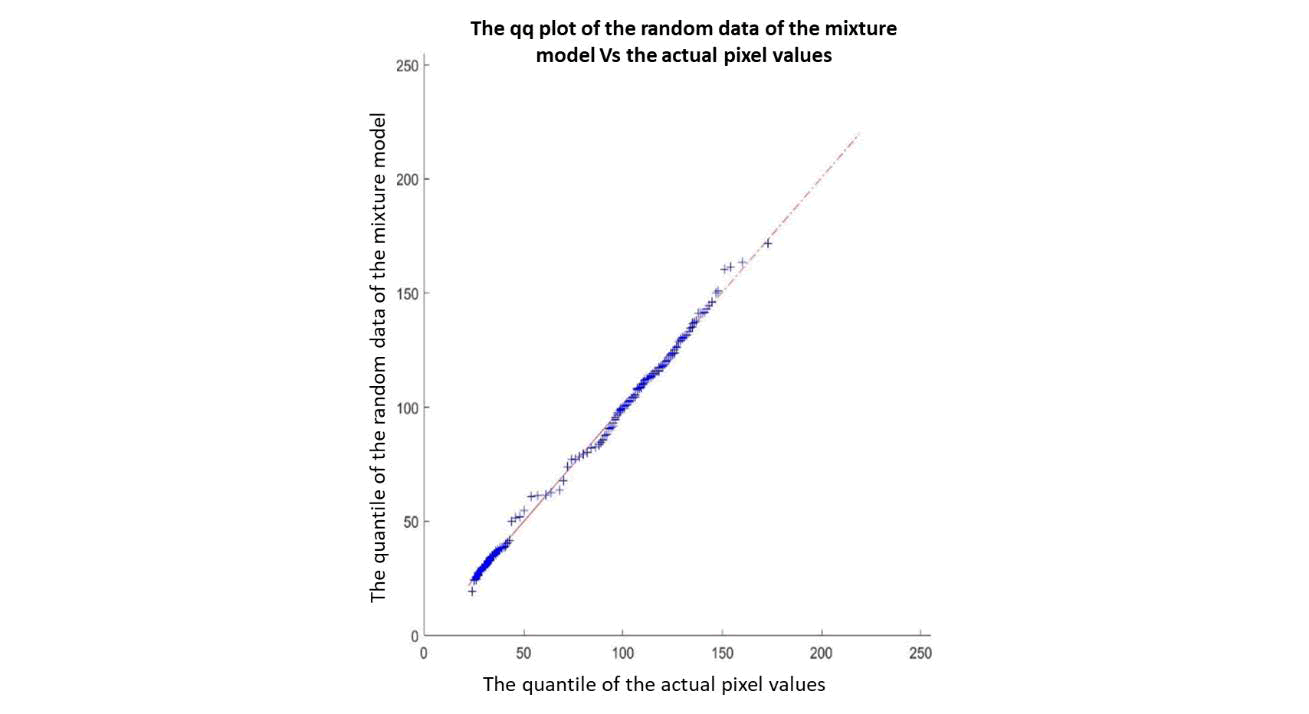
Next, a quantile-quantile plot (qq plot) was conducted to assess the fit of the previously conducted mixture model. Random data was applied to the 4-class mixture model, and then compared with the actual pixel data of the N3 image. Figure 6 shows the quantile-quantile plot of the random data of the mixture model against the actual pixel values of the N3 image. The dashed line (in red) represents the perfect case, corresponding to the samples being compared. The blue "+" sign indicates the location of the matched quantiles along the red line. From the graph, it can be seen that the distributions of the samples have almost the same shape (the "+" sign coincides with the dashed straight line). Thus, it can be concluded that the mixture model is a suitable model to be applied to the N3 image classification.
Figure 6. Quantile-quantile plot (qq plot) of random data of the mixture model against actual pixel values of image N3.
Conclusion
From the process of identifying pdf pixel values of DW MRI images of the brain, namely N1, N2, and N3, to find the probability of certain pixel values so that they can be classified into a number of optimal classes, namely 4; 3; 4 classes, respectively. In the process of classifying the N1 image, it is shown that the optimal number of classes obtained is as many as 4 classes with mean ± SD values of pixel intensity of classes 1 to 4 respectively of 1.84 ± 2.17; 9.51 ± 5.03; 83.49 ± 25.98; 236 ± 0 (range of gray scale values 0-255). In the N2 image, the optimal number of classes obtained after 12 iterations is 3 classes with a mean ± SD value of pixel intensity of classes 1 to 3 respectively of 8 ± 0; 14.11 ± 5.55; 49.28 ± 6.46. In image N3, the optimal value of 4 classes is obtained, after going through the iteration process 200 times with the mean ± SD value of pixel intensity of classes 1 to 4 successively amounting to 28.89 ± 1.81; 33.61 ± 3.71; 98.70 ± 33.27; 114.19 ± 15.15.
By applying the mixture model to the three images, the effect on image contrast has been observed from the statistical values (weight, mean and standard deviation) of each class. From the results, it can be seen that each class has a different mean value. If the difference in average pixel intensity is large (for example, in the 2nd and 3rd classes of image N1), then the two classes have a large contrast. Whereas in the two classes that have a small difference in average pixel intensity (for example in the 1st and 2nd classes of image N3), the two classes have low contrast.
In addition, the results of classifying MRI images using the ML-EM mixture model are also shown through the maximum log-likelihood function graph which corresponds to the results of the quality fit test using the minimum BIC. At the end of the research phase, the fit test of this mixture model has also been carried out on random data. Thus, it can be concluded that the mixture model is a suitable and effective model in classifying certain areas in DW MRI images of the brain.
References
- Human Memory. Brain neurons. (2024).
- WebMD. Meningioma: Causes, symptoms, and treatment. (2024).
- UCLA Health. Meningioma brain tumor. (2024).
- Demirkaya O, Asyali MH, Sahoo PK. Image processing with MATLAB: Applications in medicine and biology. Florida. (2009).
- Tsougos I. Advanced MR neuroimaging: From theory to clinical practice. Florida. (2018).
- Acampra A, Romeo V, Maurea S et al. High b-value diffusion MRI to differentiate recurrent tumors from Post-treatment changes in head and neck squamous cell carcinoma: A single center prospective study. Biomed Res Int. (2016).
[Crossref] [Google Scholar] [PubMed]
- Villalon-Reina JE, Garyfallidis E. DIPY: Brain tissue classification. GigaScience. 5 (2016).
[Crossref]
- Radiopaedia. Wide-based illumination and detection in functional near-infrared spectroscopy for enhanced seizure detection in grey matter. Normal MRI Brain. 25 (2025).
[Crossref] [Google Scholar] [PubMed]
- Redner R, Walker H. Mixture densities, maximum likelihood and the EM algorithm. SIAM Rev. 26, 195-202 (1984).