Editorial - Pharmaceutical Bioprocessing (2018) Volume 6, Issue 1
Creating cell-free protein synthesis factories
- *Corresponding Author:
- Cleo Kontoravdi
Department of Chemical Engineering
Imperial College London
SW7 2AZ, UK
E-mail: cleo.kontoravdi98@imperial.ac.uk
Abstract
Keywords
cell-free protein synthesis, synthetic biology, CHO cells, glycoproteins, multi-product facility
Introduction
The synthesis of therapeutic proteins has traditionally been dominated by cell-based expression systems. Among these, mammalian cell lines have become the favoured choice for the synthesis of biologically active proteins due to the necessity for appropriate post translational modifications (PTMs) [1]. Nearly 70% of all recombinant protein therapeutics are industrially produced in Chinese hamster ovary cells, which make them the most well established mammalian host cell line today [2].
However, working with these living cells has a considerable downside, as protein synthesis is always strongly dependent on host cell metabolism in determining the product yield and quality. At all times, process conditions are a compromise between the conditions that are essential for cell growth and viability, and the preferred conditions for the synthesis of a functional target protein [3]. Attention has hence been drawn to cell-free protein synthesis (CFPS) systems, which do not require intact host cells [4]. Cell-free systems have evolved from an analytical tool into a powerful complementary approach to cellbased production systems. Particularly, in the context of high-throughput production of protein libraries [5], and synthesis of difficult-to- express proteins, such as membrane [6] or even toxic proteins [7], cell-free synthesis have shown to offer the potential for rapid, flexible, and reliable protein production.
Design challenges of cell-free systems
“Like transformer toys, cell-free methods convert a living cell into a catalytic system with dramatically enhanced capabilities.” [8].
CFPS systems share the same principle of development: crude cell extracts are taken from cultured cell, and endogenous nucleic acids are removed. The lysate containing its transcriptional and translational machineries is then mixed with an external supply of energy sources, amino acids, and target DNA template to produce the protein of interest [9]. In the majority of systems, an orthogonal RNA polymerase (usually the T7 polymerase) is added to promote high yield RNA expression. Depending on the specific organism from which the extract was taken, each of the CFPS systems has its disadvantages and advantages in terms of product yield, protein folding, PTMs, cost efficiency, speed and ease of use [10]. Reactions can be either performed in so-called linked or coupled mode, in which transcription and translation happen separately or simultaneously in one reaction compartment [1]. Commonly, cell-free reactions are conducted in coupled batch mode due to ease of handling, scalability and higher cost efficiency [11]. However, in order to prolong reaction times and achieve higher product yields, more complex dialysis systems, known as continuous flow cell-free (CFCF) systems [12] and continuous exchange cell-free (CECF) systems [13], have emerged. These systems allow for the extended supply of energy and feeding substrates into the reaction chamber and the continuous removal of deleterious by-products from the reaction compartments [14].
One of the biggest challenges of CFPS remains the ATP regeneration system to power protein synthesis [15]. To date, costly high-energy phosphate bond donors such as creatine phosphate (CrP) are widely used. However, these not only lead to high operating costs, but also inhibit long-term protein synthesis due to fast energy depletion, deactivation of translational machinery, and production of inhibitory by-products in the reaction system [16,17]. Recent efforts have hence focused on using endogenous enzymes and glycolytic intermediates to activate the central energy metabolism in cell-free reactions. By employing a more physiological environment to the system and introducing slowly-metabolized energy substrates like fructose-1,6-bisphophate, glucose or even polysaccharides such as starch, the ATP supply could be remarkably extended [18,19,15]. While cell-free systems still lag behind traditional cell-based production of 5-10 g/L, they are continuously improving. Current protein yields now exceed 1 g of protein per L of reaction volume, and reaction scale has hit the 100-L milestone [20]; an achievement deemed impossible just over a decade ago.
Another main challenge is the ability of CFPS systems to perform human-like PTMs [21]. While the reaction environment is a variable that can be manipulated due to the open nature of the system (e.g. to promote disulphide bond formation [22], the lack of compartmentalization still poses serious problems in terms of protein folding and glycosylation, as well as product separation and downstream purification [23]. The well characterised CHO cell framework is expected to continue to play a major role in the development of novel therapeutic proteins in cell-based and CFPS systems thanks to their ability to perform proper folding and PTMs and its established safety data [24]. CHO-based extracts contain endogenous vesicle-like artefacts (microsomes), which are translocationally active and have already shown to successfully enable various PTMs [1,16]. However, to the best of our knowledge, detailed information about their specific composition and the glycosylation patterns that can be achieved is still missing. One of the key milestones thus remains the successful introduction of compartmentalization in cell-free systems to produce fully-functioning post-translationally modified protein therapeutics.
Potential applications and outlook
Cell-free methods show several unique advantages over traditional cell-based platforms. The reasons for this can be related to the direct access to the reaction network and avoidance of mechanisms that have evolved for preserving homeostasis. While cell-based expression is limited to the production of proteins that do not significantly affect the physiology of the host cell [25], CFPS platforms are potentially not subject to product limitations [26]. Cytotoxic, unstable, or insoluble proteins, such as membrane proteins, can all be expressed by CFPS systems [27,28]. In contrast to synthetic cells, except for the PURE system [29], cell-free systems are generally designed from a top-down approach and their biological machinery is not encapsulated in a phospholipid membrane. Hence, their reaction system remains completely accessible, which allows for direct monitoring, manipulation and control of the reaction environment [8]. Various interactions, such as protein-protein, protein-DNA, protein-RNA, protein-ligand etc., can be easily studied in CFPS systems offering a huge advantage for directed protein evolution [30]. The non-confined reaction compartment further enables the expansion and even reassignment of the genetic code by allowing the incorporation of unnatural amino acids into proteins [31]. Different techniques, such as pre-acylated tRNAs [32] or engineered ribosomes [33], exist to alter the characteristics of the final protein making cell-free systems a powerful tool for the generation of novel proteins for diverse biophysical and biotechnological applications [34]. In this context, it is also important to point out that CFPS systems allow protein expression from PCR-based templates without the need for time-consuming cloning procedures. Synthesis of a given target protein can therefore take place within one to two days, whereas the cell-based approach can take up to two weeks including the necessary cloning and cell transformation steps prior to protein production [4]. CFPS platforms are therefore seen as a valuable tool for high-throughput functional and structural analyses [35]. Finally, they are agnostic to the product and can be used on demand in a multi-product facility.
Looking into the future, the use of mammalian CFPS systems on an industrial level is perhaps still several years away, but it is clear that the potential to create a new class of smart micromachines that can be engineered for functional purposes is already present [36].
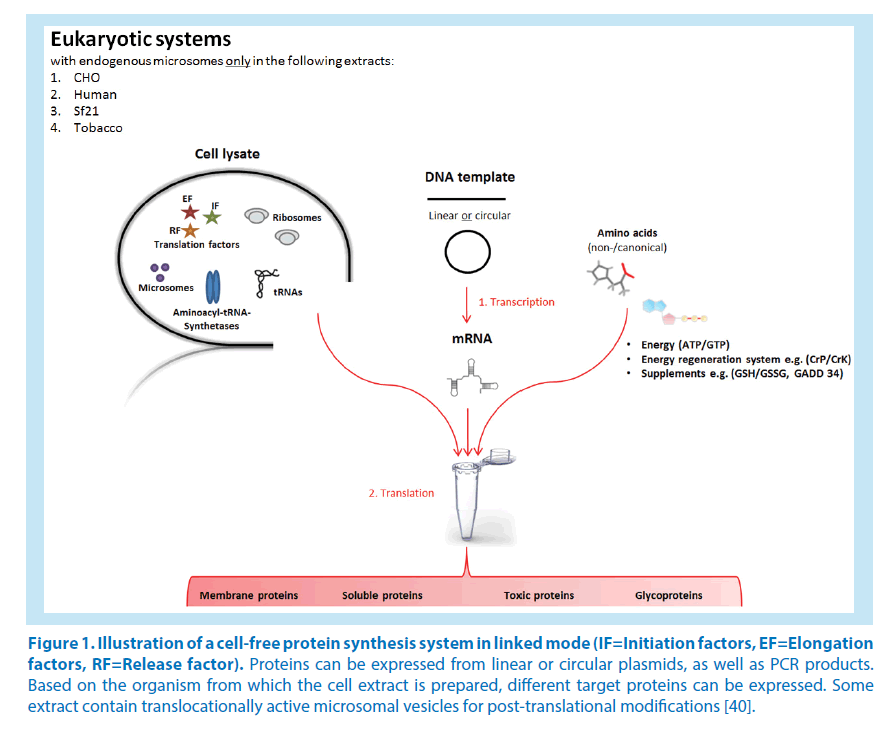
The unique advantages over traditional cell-based production will continue to be the major driving principle behind research in this area. Further growth of the field and the realization of its wider potential will, however, depend on the development of novel strategies for improved protein quality, yields and cost efficiency [37]. The introduction of compartmentalization for post-translation modifications remains crucial for achieving more complex functionality. Effective synergies with related disciplines, in particular chemical biology, and lipid membrane mechanics will help to promote the creation of distinct biochemical reaction environments within the open reaction system. For improved cost efficiency and protein yields, cell-free microfluidic reactors in continuous exchange mode have already shown promise [38]. At present, the cost of the starting reagents, in particular the energy substrates, is still too high. Novel approaches to supplement or even replace the conventional high-energy phosphate donors with more productive and sustainable ATP regeneration systems will hence play an essential role towards large-scale commercial use [39]. In addition to great advances to power CFPS with glucose and polysaccharides, such as maltodextrin and even starch even more novel ideas could be envisaged (Figure 1).
Figure 1. Illustration of a cell-free protein synthesis system in linked mode (IF=Initiation factors, EF=Elongation factors, RF=Release factor). Proteins can be expressed from linear or circular plasmids, as well as PCR products. Based on the organism from which the cell extract is prepared, different target proteins can be expressed. Some extract contain translocationally active microsomal vesicles for post-translational modifications [40].
References
- Brödel AK, Kubick S. Developing cell-free protein synthesis systems: a focus on mammalian cells. Pharm. Bioprocess. 2(4), 339-348 (2014).
- Jayapal KP, Wlaschin KF, Hu W et al. Recombinant protein therapeutics from CHO cells-20 years and counting. Chem. Eng. Prog. 103(10), 40 (2007).
- Stech M, Brodel AK, Quast RB et al. Cell-free systems: functional modules for synthetic and chemical biology. Adv. Biochem. Biotechnol. 137, 67-102 (2013).
- Carlson ED, Gan R, Hodgman CE et al. Cell-free protein synthesis: applications come of age. Biotechnol. Adv. 30(5), 1185-1194 (2012).
- Griffiths AD, Tawfik DS. Directed evolution of an extremely fast phosphotriesterase by in vitro compartmentalization. Embo. J. 22(1), 24-35 (2003).
- Katzen F, Peterson TC, Kudlicki W. Membrane protein expression: no cells required. Trends. Biotechnol. 27(8), 455-460 (2009).
- Bechlars S, Wüstenhagen DA, Drägert K et al. Cell-free synthesis of functional thermostable direct hemolysins of Vibrio parahaemolyticus. Toxicon. 76, 132-142 (2013).
- Swartz JR. Transforming biochemical engineering with cell-free biology. AIChE. J. 58, 5-13 (2012).
- Spirin AS, Swartz JR. Cell-free Protein Synthesis: Methods and Protocols, Wiley (2014).
- Braun P, LaBaer J. High throughput protein production for functional proteomics. Trends. Biotechnol. 21(9), 383-388 (2003).
- Kim DM, Swartz JR. Prolonging cell-free protein synthesis with a novel ATP regeneration system. Biotechnol. Bioengg. 66(3), 180-188 (1999).
- Spirin A, Baranov V, Ryabova L et al. A continuous cell-free translation system capable of producing polypeptides in high yield. Sci. 242(4882), 1162-1164 (1988).
- Alakhov JB, Baranov VI, Ovodov SJ et al. Method of preparing polypeptides in cell-free translation system (1995).
- Schwarz D, Klammt C, Koglin A et al. Preparative scale cell-free expression systems: new tools for the large scale preparation of integral membrane proteins for functional and structural studies. Methods. 41(4), 355-369 (2007).
- Anderson MJ, Stark JC, Hodgman CE et al. Energizing eukaryotic cell‐free protein synthesis with glucose metabolism. FEBS. Letters. 589(15), 1723-1727 (2015).
- Mikami S, Kobayashi T, Yokoyama S et al. A hybridoma-based in vitro translation system that efficiently synthesizes glycoproteins. J. Biotechnol. 127(1), 65-78 (2006).
- Kim HC, Kim DM. Methods for energizing cell-free protein synthesis. J. Biosci. Bioeng. 108(1), 1-4 (2009).
- Wang Y, Zhang YH. Cell-free protein synthesis energized by slowly-metabolized maltodextrin. BMC. Biotechnol. 9(1), 58 (2009).
- Kim HC, Kim TW, Kim DM. Prolonged production of proteins in a cell-free protein synthesis system using polymeric carbohydrates as an energy source. Process. Biochem. 46(6), 1366-1369 (2011).
- Kwon YC, Jewett MC. High-throughput preparation methods of crude extract for robust cell-free protein synthesis. Sci. Rep. 5, 8663 (2015).
- Brödel AK, Sonnabend A, Kubick S. Cell-free protein expression based on extracts from CHO cells. Biotechnol. Bioeng. 111(1), 25-36 (2014).
- Goerke AR, Swartz JR. Development of cell-free protein synthesis platforms for disulfide bonded proteins. Biotechnol. Bioengg. 99(2), 351-367 (2008).
- Stech M, Kubick S. Cell-Free synthesis meets antibody production: a review. Antibodies. 4(1), 12-33 (2015).
- Kim JY, Kim YG, Lee GM. CHO cells in biotechnology for production of recombinant proteins: current state and further potential. Appl. Microbiol. Biotechnol. 93, 917-930 (2012).
- Chrunyk BA, Evans J, Lillquist J et al. Inclusion body formation and protein stability in sequence variants of interleukin-1 beta. J. Biol. Chem. 268, 18053-18061 (1993).
- Sawasaki T, Ogasawara T, Morishita R et al. A cell-free protein synthesis system for high-throughput proteomics. Proc. Natl. Acad. Sci. USA. 99(23), 14652-14657 (2002).
- Stiege W, Erdmann VA. The potentials of the in vitro protein biosynthesis system. J. Biotechnol. 41, 81-90 (1995).
- He M, Taussig MJ. Ribosome display: cell-free protein display technology. Brief. Funct. Genomic. Prot. 1(2), 204-212 (2002).
- Shimizu Y, Inoue A, Tomari Y et al. Cell-free translation reconstituted with purified components. Nat. Biotech. 19(8), 751-755 (2001).
- Golynskiy MV, Haugner JC, Morelli A et al. In vitro evolution of enzymes. Methods. Mol. Biol. 978, 73-92 (2013).
- Hartman MCT, Josephson K, Szostak JW. Enzymatic aminoacylation of tRNA with unnatural amino acids. Proc. Nat. Acad. Sci. USA. 103(12), 4356-4361 (2006).
- Forster AC, Tan Z, Nalam MN et al. Programming peptidomimetic syntheses by translating genetic codes designed de novo. Proc. Natl. Acad. Sci. USA. 100(11), 6353-6357 (2003).
- Ohuchi M, Murakami H, Suga H. The flexizyme system: a highly flexible tRNA aminoacylation tool for the translation apparatus. Curr. Opin. Chem. Biol. 11(5), 537-542 (2007).
- Shimizu Y, Kuruma Y, Ying BW et al. Cell-free translation systems for protein engineering. FEBS. J. 273(18), 4133-4140 (2006).
- Whittaker JW. Cell-free protein synthesis: the state of the art. Biotechnol. Lett. 35(2), 143-152 (2013).
- Elani Y, Law RV, Ces O. Vesicle-based artificial cells: recent developments and prospects for drug delivery. Ther. Del. 6(5), 541-543 (2015).
- Katzen F, Chang G, Kudlicki W. The past, present and future of cell-free protein synthesis. Trends. Biotechnol. 23(3), 150-156 (2005).
- Niederholtmeyer H, Stepanova V, Maerkl SJ. Implementation of cell-free biological networks at steady state. Proc. Natl. Acad. Sci. USA. 110(40), 15985-15990 (2013).
- Caschera F, Noireaux V. A cost-effective polyphosphate-based metabolism fuels an all E. coli cell-free expression system. Metab. Engg. 27, 29-37 (2015).
- Zemella A, Thoring L, Hoffmeister C et al. Cell-Free Protein Synthesis: Pros and Cons of Prokaryotic and Eukaryotic Systems. Chem. biochem. 16(17), 2420-2431 (2015).